Native aggregation
as a cause of origin of temporary cellular structures
needed for all forms of cellular activity, signaling and
transformations
Theoretical Biology and Medical Modelling 2010, 7:19.
Laboratory of Cell Physiology, Institute of Cytology,
Russian Academy of Sciences,
Tikhoretsky Ave
E-mail: vladimir.matveev @ gmail.com
Full text [PDF]
Abstract.
According to the hypothesis explored in this paper, native aggregation is
genetically controlled (programmed) reversible aggregation that occurs when
interacting proteins form new temporary structures through highly specific
interactions. It is assumed that Anfinsen's dogma may be extended to protein
aggregation: composition and amino acid sequence determine not only the
secondary and tertiary structure of single protein, but also the structure of
protein aggregates (associates). Cell function is considered as a transition
between two states (two states model), the resting state and state of activity
(this applies to the cell as a whole and to its individual structures). In the
resting state, the key proteins are found in the following inactive forms:
natively unfolded and globular. When the cell is activated, secondary
structures appear in natively unfolded proteins
(including unfolded regions in other proteins), and globular proteins begin to
melt and their secondary structures become available for interaction with the
secondary structures of other proteins. These temporary secondary structures
provide a means for highly specific interactions between proteins. As a result,
native aggregation creates temporary structures necessary for cell activity.
"One of the principal objects of theoretical research in any department of
knowledge
is to find the point of view from which the subject appears in its greatest
simplicity."
Josiah Willard Gibbs (1839-1903)
Introduction.
To date, numerous mechanisms, signal pathways, and different factors have been
found in the cell. Researchers are naturally eager to find commonalities in the
mechanisms of cellular regulation. I would like to propose a substantial
approach to problems of cell physiology – the structural ground that produces
signals and underlies the diversity of cellular mechanisms.
The methodological basis for the proposed hypothesis results
from studies by the scientific schools of Dmitrii Nasonov [1] and Gilbert Ling
[2-6], which have gained new appreciation over the last 20-30 years owing to
advances in protein physics [7] in the study of properties of globular proteins,
their unfolding and folding, as well as the discovery of novel states of the
protein molecule: the natively unfolded and the molten globule. The key
statement for the rationale of the present paper is that the specificity of
interactions of polypeptide chains with each other (at the intra- and
inter-molecular levels) can be provided only by their secondary structures,
primarily α-helices and β-sheets.
Nasonov’s school discovered and studied a fundamental
phenomenon — the nonspecific reaction of the cell to external actions [1], while
works by Ling [5] and his followers allow the mechanisms of this phenomenon to
be understood.
The above-mentioned cell reaction has been called
nonspecific because diverse physical and chemical factors produce the same
complex of structural changes in the cell: an increase in the turbidity and
macroscopic viscosity of the cytoplasm and in the adsorption of hydrophobic
substances by cytoplasmic proteins. It is of primary importance that the same
changes also occur in the cell during its transition into the active state:
muscle contraction, action potential, enhancement of secretory activity (for
details, see [8]). Hence, from the point of view of structural changes, there is
no fundamental difference between the result of action on the cell of
hydrostatic pressure and, for instance, muscle contraction. In both cases,
proteins are aggregated.
Nasonov called the cause of these changes the stages of cell
protein denaturation, as the changes of properties of isolated proteins during
denaturation are very similar to the changes in the cytoplasm during the
nonspecific reaction. As a result, the denaturational theory of cell excitation
and damage was created [1]. The structural changes of protein denaturation were
unclear in Nasonov’s time. Nowadays, it is assumed that the denaturation is the
destruction of the tertiary and secondary structure of a protein. Below I give
two definitions, for the denaturation of natively folded (globular) proteins and
for natively unfolded proteins.
A key notion in physiology is the resting state of the cell. This is implicit in the concept of the threshold
character of the action of stimuli on the cell, which has played a historical
role in the development of physiological science. It is the threshold that is
the boundary between two states — rest and activity. But in effect, all our
knowledge about cells concerns active cells, not cells in the resting state. It
is in the active cell that variable changes occur that can be recorded. Nothing
happens in the resting cell, so there is nothing to be recorded in it.
Nevertheless, it is obvious that the resting state is the initial cell state,
the starting point for all changes occurring in the cell.
What characterizes the structural aspect of the cell in the
state of rest? It is only in Ling’s work [5] that I have found a clear answer to
this question. The answer can be interpreted as follows: if all resting cell
proteins were arranged in one line, it would turn out that most of the peptide
bonds in this superpolypeptide would be accessible to solvent (water), while
only a few would be included in secondary structures. When the cell is
activated, the ratio between the unfolded and folded areas is changed sharply to
the opposite: the proportion of peptide bonds accessible to solvent decreases
markedly, whereas the proportion included in secondary structures rises
significantly. These two extreme states of cell proteins, suggested by Ling,
provide a basis for further consideration.
If Ling’s approach is combined with Nasonov’s theory, we
obtain several interesting consequences. First of all, it is clear that proteins
with maximally unfolded structures form the structural basis of resting cells
because they are inactive, i.e., do not interact with other proteins or other
macromolecules. The situation changes when an action on the cell exceeds the
threshold: completely or partially unfolded key proteins begin to fold when new
secondary protein structures are formed. Owing to these new secondary
structures, the proteins become capable of reacting, i.e., intramolecular
aggregation (folding of individual polypeptides into globules) and
intermolecular aggregation (interaction of some proteins with others) begin. A
distinguishing feature of these aggregational processes is their absolutely
specific character, which is ensured by the amino acid composition, shape, and
size of the secondary structures. The structures appearing have physiological
meaning, so such aggregation is native and the secondary structures causing it
are centers of native aggregation. Another source of secondary structures
necessary for native aggregation is the molten globule.
The ability of cells to return to the initial state, the
state of rest, means that native aggregation is completely reversible, and the
structures appearing in the course of native aggregation are temporary and are
disassembled as soon as they cease to be necessary. Native aggregation can
involve both the whole cell and individual organelles, compartments, and
structures, and activation of proteins is of a threshold rather than a
spontaneous character.
The meaning of the proposed hypothesis of native aggregation
is that the primary cause of any functional changes in cell is the appearance,
as a result of native aggregation, of temporary structures, continually
appearing and disintegrating during the life of the cell. Since native
aggregation is initiated by external stimuli or regulatory processes and the
structures appearing have a temporary character, these structures can be called
signal structures.
Signal structures can have different properties: (i) they
can be centers of binding of ions, molecules (solutes), and proteins; (ii) they
can have enzymatic activity; (iii) they can form channels and intercellular
contacts; (iv) they can serve as matrices organizing the interactions of
molecules in synthetic and transport processes; (iv) they can serve as receptors
for signal molecules; (v) they can serve as the basis for constructing even more
complex supramolecular structures. These structures “flash” in the cell space
like signal lights, perform their role, and disappear, to appear in another
place and at another time. The meaning of the existence of the structural
“flashes” is that during transition into the active state the cell needs new
resources, functions, mechanisms, regulators, and signals. As soon as the cell
changes to the resting state, the need for these structures disappears, and they
are disassembled. Extreme examples of native aggregation are muscle contraction,
condensation of chromosomes, the appearance of the division spindle, and
interactions of ligands with receptors.
Thus, the present paper will consider the meaning and
significance of native aggregation as the universal structural basis of the
active cell. The basis of pathological states is the inability of the cell to
return to the resting state and errors in the formation of signal structures.
The presentation of native aggregation is based on three pillars: (i) reversible
protein aggregation is a structural basis of cell activity (Nasonov's School);
(ii) the operation of the living cell or its individual structures can be
regarded as a repetitive sequence of transitions between two states (active and
resting), a key role in which belongs to natively unfolded proteins (Ling's
approach); (iii) the specificity of interactions of separate parts of a single
polypeptide chain with each other (folding) or the interaction of separate
polypeptide chains among themselves (self-assembly, aggregation) can be provided
only by protein secondary structures.
The goal of this paper is the enunciation of principles,
rather than a review of facts corresponding to these principles.
Native
aggregation in retrospective. The best-studied nonspecific
response of cells to external actions might possibly be the response to
fixatives. For a long time in the history of science, cells were considered optically
empty structures by researchers. The appearance of methods of fixation and
staining wrought a revolution in cytology, as these approaches opened to the
researchers’ sight numerous cell structures whose existence had not even been
suspected. After a period of euphoria, doubts were cast: were these structures
real or were they the results of fixation, denaturation
of the cell’s native substance?
The danger of serious errors when artifacts of fixation might be considered
real structures became a subject of general attention after 1899 (see [9], Ch.
1 for details), when coagulation of homogenous protein solutions was shown to
lead to the appearance of structures quite similar to those observed in fixed
cell preparations (see [10], Fig. twenty-four). The shape of such artificial
structures depended on the chemical nature of the fixative, its concentration,
the protein concentration in solution, the temperature, and other conditions.
This brought about an obvious crisis in the study of cell morphology.
However, other things were also obvious. In the optically empty part of the
cell, visible structures could appear not only during fixation, but also during
the transition of the cell to the active state. Comparative observations on
fixed preparations and living cells showed that where the structure appeared in
vivo, it was also observed in a fixed preparation. The obvious resemblance
between native structures and the structures obtained as a result of fixation
gave grounds for considering that several cell structures are formed not only
at fixation, but also during activation of some particular cell fraction, when
new structures absent in the resting cell are formed by self-assembly (see [9],
Ch. 1 for details).
This discussion has led to the rather important conclusion that despite the
dangers of producing artifacts, another thing is beyond doubt: in the process
of aggregation, the denatured cell proteins interact with each other not
chaotically, but regularly, in accordance with a certain plan (this is what I
call native aggregation). The laws of this interaction lead to the formation of
temporary structures necessary for the cell to function under new conditions.
During fixation and dehydration, this process initially occurs “as it should”
(the self-assembly of real cellular structures takes place), but it goes too
far when the process of making the preparation is completed, when aggregation
becomes irreversible and the structure appearing as a result of aggregation
becomes a “corpse”. If the interaction of proteins during aggregation had been
chaotic, we would still know little about cell structure.
The course of native aggregation seems to be determined by the non-homogeneity
of the content of the resting cell; it has structure that is invisible under
the light microscope, but reveals itself at the onset of native aggregation.
The role of structure guiding native aggregation may be played, for instance,
by Porter’s “microtrabecular lattice” [11], which can be envisaged «…as that
which is in the background of all the visible membranous organelles and all the
visible elements of the cytoskeleton; e.g., that which has been invisible up
until now and which we wish to “see” microscopically» [12]. Such a lattice
might act as the center of “crystallization” or the center of “attachment” of
aggregating proteins. However, this is merely an example that I cite for
clarity. The centers of crystallization can also comprise the most sensitive
proteins that are the first to respond by conformational alterations to changes
in the medium and become aggregation-competent. In any event, as a result of
native aggregation, the hidden structures become visible under the microscope.
Fulton [13], a convinced Porter devotee, moved even further: she put forward a
point of view that “the cytoplasm is so compact that it is only occasionally
more open than a crystal”. Sufficient data have probably accumulated in the
literature to establish that the content of a cell is to be considered a
structured system that guides native aggregation into the required course. As
one example, one can indicate the data of Baló-Banga
et al. [14]: the birefringence of lymphocyte nuclei was enhanced after fixation
with ethanol, i.e., correct fixation leads to the appearance of new, more
ordered structures. However, especially interesting are the cases when native
aggregation, as I call it, takes place in the process of normal cell
functioning. Thus, in the same work by Baló-Banga
et al. [14], activation of lymphocytes by specific antigens or haptens was
shown to lead to a significant enhancement of nuclear birefringence. The same
phenomenon was also observed in the case of activation of peripheral blood
lymphocytes with allergens in drug-allergic patients [15].
If the factor affecting the cell becomes more intense, its activating
effect will be replaced by a damaging one. Thus, the studies of Inners and
Bendet [16] on thermal DNA denaturation in bacteriophage T2 and spermatozoa
[17] showed that during irreversible denaturation of structures their capacity
for birefringence is lost. Such data indicate that under certain conditions,
the actions of heat, organic solvents, etc. on cells produce not native
aggregation, but destruction, disorganization of intracellular structure; in
other words, destruction of structure can follow native aggregation.
Unfortunately, there is a marked tendency in the literature towards rough
alterations in the structure of the cytoplasm and organelles, because they are
easier to study.
Thus, the retrospective considered shows that when adequate methods of study
are used, native (programmed) protein aggregation leading to self-assembly of
various cell structures is the usual phenomenon of cell life. An example of
this is the universal reaction of the living cell [8].
Universal
reaction of the living cell and native aggregation. Why does
native aggregation not occur in cells in the resting state but begins only on
activation (for instance, muscle contraction, action potential) or damage? To
answer this question, let us return to Nasonov’s denaturation theory [1].
According to this theory, excitation of the cell takes place only when its
proteins are subjected to the initial stages of denaturation.
Mirsky seems to have been the first to pay attention to the similarity between
changes in active cytoplasm and the denaturation of isolated proteins [18].
Mirsky came to the conclusion that denaturational protein changes appear when
an egg cell is fertilized [19] and during photoreception [20]. This is what he
says about it in the latter of the above-cited works: "...There is
evidence indicating that light denatures a conjugated protein, visual purple,
and that denaturation reverses in the dark." However, his studies in this
direction were not systematic.
Nasonov and his followers studied the effects of quite different factors
(chemical substances, pH, hydrostatic pressure, mechanical action) on cells of
different types. As a result, a regularity was revealed: regardless of the
character of the action and the type of cell, the response reaction represented
a monotypic (nonspecific) complex of synchronous changes. These changes were of
two-phase character: macroscopic viscosity first decreased, then rose; binding
of vital dyes by cell structures (under conditions of diffuse equilibrium)
first decreased, then increased; in the first phase of the reaction the
cytoplasm became clear, in the second phase it became turbid. Other parameters
(see [8] for review) were also studied. The first phase of this reaction is not
related to the subject of the present paper, as it is a variation of the
resting state. Of interest to us is the second phase, whose structural basis is
protein aggregation (Fig. 1). It is this phase that is the phase of activation
of cell functions [1].
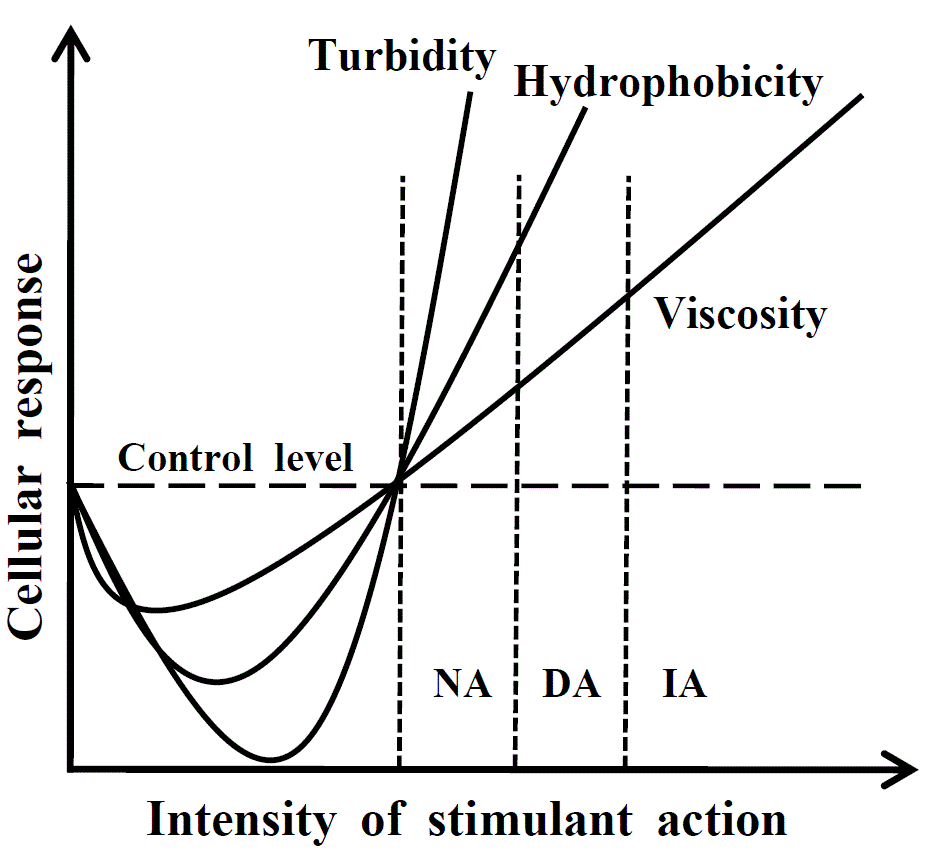
Fig. 1. Response reaction of cell
depending on strength of external action (scheme). On reaching the threshold,
the first phase of the cellular response begins; during this phase the cell
becomes more transparent, while hydrophobicity and macroscopic viscosity
decrease. Then the second phase of the cellular response begins, during which
all parameters measured significantly exceed the control level (in this case,
the control level means the cell resting state).
NA, native aggregation when necessary cellular functions and signaling pathways
are activated; DA, damaging aggregation when signals for apoptosis, cancer
transformation or other pathological cellular states may be generated; IA,
irreversible aggregation leading to cell death. See [8] for details.
This second phase was called the phase
of excitation and damage by Nasonov’s school. Substantial changes in the cell
in this phase are remarkably reminiscent of denaturation of isolated proteins;
therefore, Nasonov called his theory explaining the cell response reaction the
denaturational theory of excitation and damage. According to this theory, the
initial stages of denaturational changes, when they still are reversible,
underlie cell excitation (activation of secretory function, muscle contraction,
action potential, etc.). More profound protein changes lead to disturbances of
normal cell functioning, but may still be reversible. Then, with further
development of damage, denaturational changes become irreversible and the cell
dies.
The peculiarity of the cellular reaction discovered and studied by Nasonov’s
school was its nonspecific character: whatever the action on the cell was, its
proteins were aggregated (as in fixation); any cellular activity was also
accompanied by protein aggregation (this is especially well seen in the case of
muscle contraction). The behavior of isolated proteins during denaturation was
the same: any denaturing agent caused their aggregation (except for
denaturation under non-physiological conditions, e.g. denaturation by
concentrated solutions of urea).
In this universalism of the cellular response, a puzzle was hidden, but
in an era concerned with specific interactions, nonspecific phenomena drew no
attention. Nevertheless, it is obvious that the nonspecific cellular reaction
discovered by Nasonov’s school is a fundamental natural phenomenon – like cell
division or carcinogenesis. Attention to it is justified because the phenomena
of nature, unlike theories, cannot be erroneous.
The nonspecific character of the cellular reaction considered is a superficial
impression. Death is also a nonspecific phenomenon, but the processes leading
to it are characterized by diversity and can be extremely specific. In exactly
the same way, aggregation of proteins can be based on specific interactions. If
we deny the existence of specific mechanisms in cell protein aggregation, we
will not be able to understand why cell stress initiates such processes as
proliferation, differentiation, senescence, apoptosis, necrosis, or mitotic
cell death [21]. On the other hand, it is obvious that with all the specificity
of interactions leading to protein aggregation, the cellular reaction looks
nonspecific because any aggregation, whether specific or nonspecific, ends in
the formation of protein complexes. Therefore, it is more correct to focus not
on the nonspecificity, but on the universality of the complex of structural and
functional cellular changes studied by Nasonov’s school. That is why I have
proposed to name this typical cellular response a universal reaction of the living
cell or protoreaction, because there are grounds to consider it the most
ancient type of cellular reaction to external actions [8].
Thus, it is the denaturation of proteins that makes these polymers
active. Their activity arises from the fact that only denatured proteins begin
to interact with each other. This interaction seems to be specific and regular;
native aggregation results in new structures that are absent in the resting
state and have physiological meaning for the active state. In other words,
denaturational changes make proteins reaction-capable.
While these changes are reversible, the cell is able to disassemble the
temporary structures formed and to return to the initial state – the resting
state. When damage ensues, when protein aggregation becomes too extensive or
irreversible, pathological changes appear in the cell and can lead to its
death. The threshold character of the cellular reaction means that the resting
state and the active state are different thermodynamic states of the system,
which are separated by an energy barrier; this relates not only to the cell as
a whole, but also to its individual components [5].
Now the time has come to ask: what makes protein aggregation specific?
The answer to this question is provided by the physics of proteins. It has been
established that the correct folding of a polypeptide to a globule, like the
unique structure of the globule itself, is provided by specific interactions
between protein secondary
structures [7]. Let us consider a structure such as an α-helix. It interacts with
other secondary structures via its surface. The surface of the α-helix is “encrusted” with
polar (hydrophilic) and non-polar (hydrophobic) groups. Taken individually,
these groups are capable only of nonspecific interactions, but the secondary
structures confer a specific character on these interactions. This is their
biological meaning. Indeed, depending on the amino acid composition, the
topography of hydrophobic groups on the surface of an α-helix can vary strongly. If two α-helices have complementary topographies
of hydrophobic amino acid residues, such secondary structures will “recognize”
each other and associate to form a hydrophobic nucleus (the principle of “key”-to-“lock” correspondence works here, too). Owing to the
same topographic
factor, polar groups can form on the secondary structure surface a “landscape”
complementary to the nucleic acid surface. To provide specificity of
interaction by a unique distribution of protein functional groups on the
surface is the main purpose of all protein secondary structures. The principle
of structural complementarity has a universal physical basis and is realized
not only in intraprotein interactions (in the globular proteins formed and in
the process of their folding), but also in interprotein interactions (native
aggregation) including protein-nucleic acid interactions.
When an action on a cell or cell structure exceeds the threshold, (i) formation
of secondary structures begins in natively unfolded proteins (or unfolded regions
of proteins), while (ii) secondary structures of molten globules start to
become accessible for interaction with secondary structures of other proteins
and with nucleic acids. Such secondary structures induced by the external
action are centers (sites)
of native aggregation. Thus, the first event in the activated cell
is the appearance of new secondary structures able to interact selectively with
each other to form tertiary, quaternary, etc. structures. Proteins whose
secondary structures appear under such circumstances lose their previous
inertia and become reaction-capable.
The proposed approach to understanding the mechanisms of cellular reactions
poses the question of native and denatured protein states in a new way. In the
native state the key cell proteins are inert, non-reaction-capable; they do not
interact with each other or with other biopolymers. Loss of the state of
inertia is denaturation. On denaturation of the unfolded polypeptide chains the
secondary structures appear, whereas on denaturation of molten globules their
secondary structures are modified and “float up” to the surface from the
hydrophobic nucleus. In both cases the secondary structures are ready to
interact. In other words, two extreme protein states can be identified: the completely
folded (the globular protein) and the completely unfolded states. Between these
inactive (native) states, numerous intermediate, active forms can exist; it is
these forms that provide native aggregation. Thus, in proteins, only two states
are inactive (they are native states). In all other cases they are active, as
manifest in the capacity for native aggregation.
The proposed mechanism of native aggregation explains the increase of volume of
the cellular hydrophobic phase during the protoreaction [8] and the structural
changes in the universal reaction of the living cell [1]. When secondary
structures form, the polar groups of peptide bonds break contact with water and
form hydrogen bonds with each other. For this reason alone the hydrophobicity of
a polypeptide with secondary structures is higher than in the unfolded
polypeptide-precursor. The volume of the hydrophobic phase increases even more
when the secondary structures fuse to form hydrophobic domains (nuclei). The
second reason why the volume of the cell hydrophobic phase increases further is
the appearance of molten globules. In native globular proteins the hydrophobic
nucleus is a solid body with a comparatively small surface interacting weakly
with hydrophobic substances (therefore, the cell in the resting state is
hydrophilic). On melting, the hydrophobic nucleus ceases to be a solid body
([7], Lecture 17); its constituent elements become much more mobile relative to
each other, and the nucleus loosens and becomes accessible to water and to
substances dissolved in it (surface hydrophobic contacts increase). If the
solution contains hydrophobic compounds, it becomes possible for them to
penetrate into the molten globule nucleus and become concentrated in this
hydrophobic phase.
Proteins in the excited state are capable not only of new intramolecular
interactions, but also of interaction with other proteins. Protein physics
offers no prohibitions on this point. Native aggregation (formation of specific
aggregates) explains the increase of cell turbidity and of macroscopic
viscosity of the cytoplasm and nucleus. Thus, the observed changes during the
protoreaction are given a simple explanation based on data from protein physics
[7].
In this section, significant attention was paid to the cell in the
resting state. Let us now consider it in greater detail.
What
is the resting state of the living cell? To study any process,
it is important to identify a starting point. For instance, it would have been
impossible to understand the mechanism of muscle contraction without the
concept of the resting state of the contractile apparatus. Based on the
experience of classical physiology, it is necessary to accept that the concept
of the resting state of cell (as well as of its individual parts) is of great
importance for understanding the mechanisms of activation. Here we return again
to the issue of the structure of the resting cell. The fact that such a cell,
unlike an activated one, is almost completely transparent, indicates a
negligible amount of protein aggregate. Also, the resting cell is hydrophilic,
as under conditions of diffusional equilibrium it does not bind vital dyes [1],
which are hydrophobic [8]. These essential peculiarities of the resting cell
are to be explained by its structure.
Ling [22] was the first to suggest that the structure of the resting cell is
determined by natively unfolded proteins. This concept was finally formulated
by 1965 [23], while a summary of the development of this way of thinking was
published a decade later [6]. The most important argument in favor of this
point of view is the identity of the equilibrium
distribution of substances between the cell and the medium on the one hand, and
between the model systems and the medium on the other. The model systems
studied include cellophane dialysis bags filled with concentrated solutions of
hydrophilic and electrically neutral linear
polymers, all of whose chain links are accessible to water. The distribution
law, i.e., dependence of equilibrium distribution of substances on their
concentration in the medium, is the same for the model systems and for the
living cell. Since the distribution of substances was studied under conditions
of diffusional equilibrium, this result means that the key physicochemical
factor determining the character of the distribution is identical in the models
and the cell, and is provided by unfolded biopolymers. It seems obvious that of
all cell polymers, only proteins – the most massive cell polymers – can
possibly fulfill this role [23].
What is this factor? Both cells and models have a common peculiarity: if
the solution component studied is not absorbed on a polymer within the system,
its equilibrium concentration in the internal medium is always lower than in
the external solution. Model systems, owing to their simplicity, allow this
phenomenon to be understood: it is because substances are less thoroughly
dissolved in the system water than in the water of the external medium. Physics
provides the only possible explanation for this difference: water in the cell
and in the model systems is more ordered than bulk water; therefore, insertion
of a molecule of solute with more rigid bonds into the solvent is not
energetically advantageous, so solutes are displaced (excluded) from the
system. But why is water ordered in the presence of linear polymers? The
obvious explanation is provided by model systems comprising nothing but
polymer, water, and dissolved substance: if water is absorbed by the regularly
repeated polymer links, the water itself is ordered in the space (multilayer
adsorption). Also, in the absorbed water molecules, the electrical properties
are different.
In spite of the wide diversity of proteins, they all have absolutely identical
polypeptide backbones; differences between proteins are due only to the side
chains. The polypeptide backbone of all proteins comprises a regular
alternation of positive (NH) and negative (CO) charges in the peptide bonds;
the distance between these groups turns out to be comparable with the size of a
water molecule and with the length of the hydrogen bonds between them. In other
words, the disposition of these dipoles along the polypeptide backbone is
complementary to water structure. Another peculiarity of the peptide bond
groups is that they form hydrogen bonds either with each other (in the
secondary structures) or with water (in the unfolded regions of the polypeptide
chain) ([7], Lecture 4). However, the question arises – why does the
interaction of water with the functional groups of peptide bonds change its
properties so markedly? To answer this question, let us address the properties
of electric dipoles.
An important property of dipolar molecules is that their dipole moment is not
constant, but depends on their interaction with other dipoles [24]. Example:
the dipole moment of water in the gaseous phase is equal to 1.85 D, while in
the liquid phase it is 2.9 D. Hence, the interaction of water molecules with
each other leads to their mutual polarization – an enhancement of their own
dipole moment by 60% [25]. But what if the water molecule interacts with a
stronger dipole than itself? The dipole moment of a peptide group is 3.5 D
[26]. If water interacts with these, stronger, dipoles, its molecules will be
polarized to a greater degree and their hydrogen bonds with other molecules
will become stronger. The enhancement of hydrogen bonds makes the first
adsorptional layer stable and able to attract and to bind more and more new
free water molecules, forming more and more new adsorbed layers. Thereby,
stronger dipoles on the adsorbing surface are the key prerequisites for the
multilayer adsorption of polar molecules.
Owing to the enhancement of hydrogen bonds in the multilevel adsorbed water
layer, penetration of other molecules into it (including water itself) becomes
energetically non-advantageous, because it requires breakdown of the
intermolecular hydrogen bonds in the layer, which are stronger than in the
voluminous (bulk) phase. This explains why bound water is a poor solvent
compared with the phase in which water molecules interact only with each other.
For this thermodynamic reason, the concentration of any substance in the
absorbed phase always will be lower than in the liquid phase.
However, all begins to change if the unfolded polypeptide absorbing water
begins to fold with formation of secondary structures. In this process, peptide
groups cease to form hydrogen bonds with water and form them between each
other. The previously bound water is desorbed and acquires the properties of
voluminous (bulk) solvent [6, 23, 27]. There is convincing experimental
evidence to substantiate this point of view about the interaction of
polypeptides and other hydrophilic polymers with water [28, 29].
But what is the role of globular proteins? It is these compounds that
are the second important component of the cell in the resting state. They are
the best-studied type of proteins, performing structural and enzymatic
functions. Their solid core is inaccessible to water, while polypeptide chains
containing no secondary structures are not sufficiently expanded to affect the
state of the intracellular water fundamentally [5].
Thus, in the resting state, the physical properties of the cell protein
matrix are determined by partially or completely unfolded proteins and by globular
proteins (of course, the latter include complex proteins with several globular
domains). In the context of the present paper, such proteins can meaningfully
be called native. The structural and functional peculiarities of the cell in
the resting state are determined by unfolded proteins [5].
The question remains as to why the resting state of the cell is
relatively stable and can exist for an indefinite period. Ling believes this is
accounted for by the stabilizing effect on unfolded proteins of various ligands
bound to native unfolded proteins: ions, low-molecular organic compounds,
hormones, etc. According to Ling, the most important ligand of proteins in the
resting state is ATP [30]. If some action leads to splitting of ATP or to
dissociation of other rest-making ligands, this leads to folding of the
natively unfolded protein; secondary structures appear and make the polypeptide
reaction-capable. Native aggregation begins, in the course of which signaling
structures are formed. Natively unfolded proteins seem to be the most sensitive
elements of the resting cell, as their folded state is economically
advantageous, because when the water is desorbed the entropy of the system
increases (water is the most abundant cell component). Also, the rest-making ligands
are not firmly bound to natively unfolded proteins, as the bonds are
non-covalent, while ATP can be split enzymatically. As a result, individual
cell components or the entire cell appear as a system in which the structural
content of life activity is the reversible transition from the resting state
into the activated (excitatory) state provided by the reversible transition of
proteins from the resting (native) into the activated (non-native) state.
Principles
of native aggregation. From the point of view of the proposed
approach, reactions of the cell to external actions, various forms of cellular
activity (metabolism, division, muscle contraction, secretion, intracellular
signaling, etc.) as well as pathological states are considered on the basis of
the following statements and principles.
Native aggregation is a specific interaction of proteins with each other,
realized by interaction between the secondary structures of the aggregating
proteins. If the reaction-capable secondary structures are absent or
inaccessible for interaction, native aggregation is impossible.
The cell is considered as a system that can have only two states: the resting
state and the active (excitatory) state. The same principle is true for any
cell organelle, structure or protein molecule. For clarity, a parallel can be
presented: the excitable membrane in a state of rest or excitation.
Functionally important cell proteins in the resting state are present in one of
two states: either unfolded (completely or partly – natively unfolded proteins)
or folded to the protein globule state or any other form in which secondary
structures are inaccessible for interaction with other proteins. These states
are considered the resting states of protein molecules or as their native states.
Proteins in the native state are stabilized by rest-making ligands and/or
chemical modifications, for instance, by phosphorylation/dephosphorylation.
According to Ling [30], the resting state of an unfolded protein is maintained
by its bound ATP, ions (Na+, K+, Ca2+),
molecules of bound water, hormones (for instance, insulin), and any other
significant interactions. For instance, analysis of amino acid sequences in the
regions surrounding known phosphorylation sites reveals a strong propensity
towards adoption of a natively unfolded conformation [31]. Disruption of bonds
with ligands (for instance, breakdown of ATP) leads to activation of the
protein, its transition from the resting to the active state; the same result
is produced by a decrease in the cell ATP content below the critical level.
Ling’s concept of the capability of small molecules for specific binding with
natively unfolded proteins is confirmed, for instance, in the work by
Mukhopadhyay et al. [32].
On activation of the cell by external actions, intracellular factors,
and signals of different nature (including chemical modification), a new
protein fraction appears — activated proteins with newly formed secondary
structures that were absent in the resting state (Fig. 2). These new structures
appear on the folding of natively unfolded proteins and on melting of protein
globules. They include α-helices,
β-sheets, and other secondary structure variants. The
secondary structures of activated proteins are new “valences” necessary for new
interactions – intramolecular (folding) and intermolecular (native
aggregation). In the case of large proteins, the secondary structures can form
hydrophobic sites on the protein surface, which interact specifically with
similar (complementary) structures on the surfaces of other proteins.
The natively unfolded proteins can be called excitable proteins. Their
transition to the excitatory state triggers native aggregation.
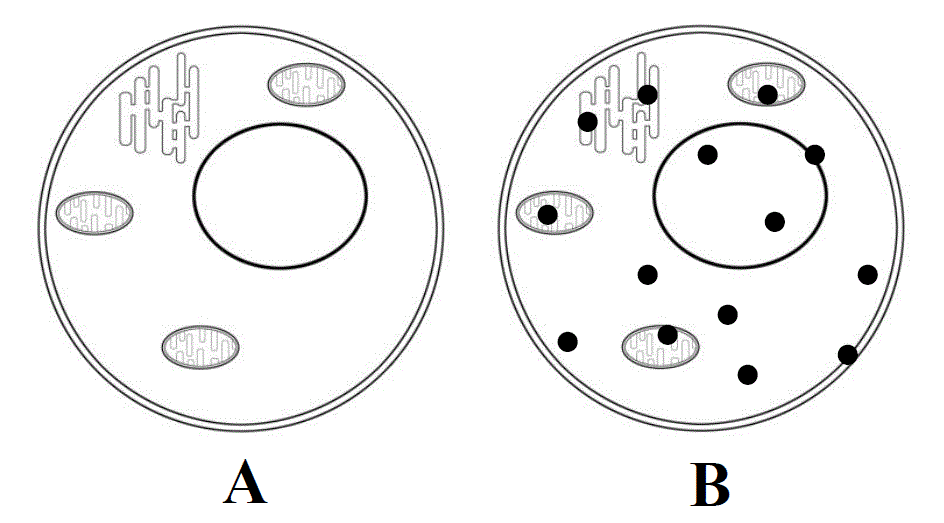
Fig. 2. Two main cell states. A - cell in the resting state, optically transparent. B - activated cell, in the cytoplasm and organelles of which native aggregation centers (closed cycles) appear - the reaction-capable secondary structures of activated proteins, owing to which native aggregation of cell proteins begins.
If on the unfolding of a globule (or a
globular domain) no molten globule intermediate is formed, while the protein
cooperatively assumes the completely unfolded configuration at once, this means
it is inactivated, as a protein without reaction-capable secondary structures
is incapable of aggregation. The molten globule may be inactivated in two ways:
by transition back to the well-folded conformation (when secondary structures
are hidden from interaction) or by unfolding of the molten globules until a
completely unfolded conformation is reached, devoid of the secondary structures
that are key to native aggregation.
The secondary structures in activated proteins play the role of centers of
native aggregation. It is these structures that provide for specific
interactions of activated proteins with each other (native aggregation) to form
new structures that have signaling and functional significance for the active
cell. Native aggregation is determined by the same forces and interactions that
are involved in the well-studied folding of unfolded polypeptides to globules.
This rule is followed: if there are secondary structures capable of specific
interaction, there is native aggregation; if there are no such structures or
they are inaccessible, there is no native aggregation.
If an action on a protein increases the number of amino acid residues included
in its secondary structures, that protein is activated and the signal pathways
in which it participates are open. If the protein is unfolded and the portion
of amino acid residues in secondary structures decreases, it undergoes
transition to the inactive state, is relaxed, while the signal pathway(s) in
which it participate(s) is/are blocked. On the transition of proteins
participating in native aggregation to the native state, native aggregates are
destroyed and individual structures and the cell as a whole transit to the resting
state.
The temporary structures appearing as a result of native aggregation perform
diverse functions. They may be centers of specific adsorption (binding) of
various ions and molecules including signal factors and proteins, i.e., can
perform the functions of receptors. They may have the enzymatic activity
necessary for performing specific functions and may serve as centers of
formation of even more complex supramolecular structures.
Only the secondary protein structures are able to provide for specificity
(selectivity) in the interaction of proteins with others, as they provide the
specificity of interactions necessary for correct folding of the polypeptide
chain to a globule (the folding of polypeptide to native globule can be
considered as intramolecular native aggregation). Each secondary structure has
a unique topology of polar and hydrophobic groups on its surface. Secondary
structures form stable complexes with each other or with sites on nucleic acids
only if their surfaces are complementary to each other, as the key is
complementary to the lock.
Native aggregation is determined genetically to the same extent as protein
structure because it is determined by the same factors that determine all
levels of organization of the individual protein molecule beginning with the
primary sequence. Secondary structures of activated (excited) proteins will
interact with other excited proteins not chaotically, but in accordance with
the genetic program. As a result of native aggregation, those structures and
corresponding functions will appear that are necessary to the cell here and
now: action potential, channels on the cell surface, in the cytoplasm and
nucleus, cytoskeleton, movement of cytoplasmic sites, cell division, apoptosis.
Errors in native aggregation that appear during a prolonged state of cell
excitation (for instance, chronic inflammation) and on damage lead to various
forms of cellular pathology: conformational diseases, necrosis, carcinogenesis.
All the differences between the excited cell and the cell in the resting state
are the direct or indirect results of native protein aggregation.
Native
aggregation in action. Since practically any change in the cell
can be considered a result of native aggregation, I will focus on only a few
examples. The aim of this section is to show how the principles of native
aggregation work in the analysis of particular phenomena.
I will begin with the natively unfolded proteins, the physical basis of the
resting state. According to Dunker et al. [33], the first data about natively
unfolded regions in proteins appeared in 1978, i.e., 26 years after Ling [22]
had first suggested their existence. Until the discovery of natively unfolded
proteins, the dominant notion was that the whole diversity of cell functions is
due only to proteins with 3D structure. Natively unfolded proteins were not
compatible with this notion and it was not clear whether they performed any
function at all. Subsequently it was found that more than 35-51% of eukaryotic
proteins had unfolded regions longer than 50 consecutive amino acid residues,
which is significantly higher than in prokaryotes [34, 35].
When it became clear that natively unfolded proteins played an important role,
Dunker et al. [33] proposed to widen the notion of functional protein types in
the cell: to proteins with 3D structure, they added molten globules and
proteins with unfolded conformations. Uversky [36] proposed to supplement this
list with a fourth, relatively stable protein conformation - the premolten
globule, which might be called the boiling globule, as in the coordinates of
the unfolding reaction it follows the globule and molten globule and precedes
the completely unfolded conformation. The rationale of this proposal is that
all four protein states are thermodynamically stable, although to different
degrees.
In the opinion of Dunker et al., transitions between different phasic states
continually take place in the cell. This is so, indeed; however, the statement
needs clarification. Let us recollect that the first ideas about the molten
globule and unfolded protein conformation were obtained by studying protein
denaturation in vitro and then they were extrapolated to the cell. Nowadays we
know that globule melting is a phase transition that fits the "all-or-nothing"
law and has a threshold, for instance, a temperature threshold [7]. This means
that several similar molecules under identical conditions will be in the same
phase state: either globule, or molten globule, or the unfolded conformation.
Within such a population, uninterrupted and asynchronous protein transitions
from one phase state to another cannot take place. However, molecules of the
same protein located in different microenvironments can be in different phase
states, but the state may also be identical for all proteins of the same
(given) population. As a result, we find that this (some) protein can indeed be
in different phase states in this cell, but only if its molecules are located
in different parts of the cell with different microenvironmental conditions.
Another specification is also to be made. According to the hypothesis of native
aggregation, there are only two basic protein states in the resting cell:
globules (here, proteins composed of two and more globular domains may be
included) and the natively unfolded state. Other transitional states appear in
the cell temporarily. They appear on reaching the threshold, when some factor
in the medium begins to produce a moderately (gentle) denaturing action. Then a
globular protein is melted, after which it unfolds (if the strength of the
action keeps rising), while natively unfolded proteins begin to fold. The
differences between the main states are fundamental: the globular conformation
is stabilized mainly by hydrophobic interactions, the natively unfolded one by
ATP and other ligands. As soon as the medium conditions return to normal, the
excited proteins are relaxed and the system returns to its main state - the
resting state.
Since native aggregation results in the appearance of signaling and regulatory
structures, it is obvious that as biological organization becomes more
complicated during evolution, more and more novel mechanisms of regulation of
the active cell are needed. This need is realized with the aid of new natively
unfolded proteins and, accordingly, of new transitory conformations appearing
as they fold.
In the literature, the mechanism of interaction of natively unfolded proteins
with protein targets has been widely discussed. Most commonly, four stages of
such interaction are identified: (i) random collision of natively unfolded
protein with target; (ii) weak, nonspecific interaction of natively unfolded
protein with target; (iii) formation of secondary structures in natively
unfolded protein; (iv) owing to these nascent secondary structures, a firm
complex of the natively unfolded protein with the protein-target is formed [37,
38].
In terms of the hypothesis of native aggregation, this scheme looks
unconvincing. Indeed, it is hard to imagine a mechanism (for instance, the mechanism
of muscle contraction) or a process in the living cell working on the basis of
random collisions. First, if natively unfolded proteins and their targets
collide randomly, it means that they are diffusing freely in the cytoplasm or
nucleus, i.e., we are dealing with a Brownian mechanism of regulation. Second,
if the first stage of interaction of the natively unfolded protein with the
target is accepted as nonspecific, this will mean that the number of
interactions of the diffusing natively unfolded protein will greatly exceed the
number of interactions necessary for the act of regulation. Under such
conditions, the correct regulatory response looks more random than regular.
From the point of view of native aggregation, these events appear differently.
The available experimental data indicate that natively unfolded proteins are
organized in clusters and oriented in space mainly in parallel to each other
([5], Ch. 11), while the protein concentration in the cytoplasm reaches 200-400
mg/mL [39]. Thus, under conditions of crowding, when the space between protein
molecules is not large and is filled with bound water ([5], Ch. 11), it is
difficult to imagine diffusion of free proteins. According to Ling, the protein
matrix of the resting cell is not chaotic, but structured. In terms of the
hypothesis of native aggregation this means that the program of protein-protein
interactions is responsible for the spatial distribution of the key matrix
elements (for instance in the contractile apparatus). Natively unfolded protein
does not diffuse in anticipation of a random hit to the target. The target is
relatively immobile and is located nearby. In the resting state they do not
interact with each other, as they are in the inactive (native) state, i.e., do
not have reaction-capable secondary structures.
If secondary structures are formed in the natively unfolded proteins during any
collision with other proteins, this will also become a random event and the
interactions of secondary structures with each other will not be amenable to
any logic. For this reason, random, nonspecific interactions are to be
eliminated from the mechanism of functioning of natively unfolded proteins. To
prevent random interactions from causing excitation of the natively unfolded
state, such proteins must be sufficiently stable. According to the proposed
approach, the natively unfolded proteins are stabilized by various ligands
depending on their property, location, and function [6].
The fourth stage of interaction with the target is also problematic
because the activated native protein will interact, in my opinion, only with
activated protein-target (with its active secondary structures). Native
globular proteins (or globular domains in large proteins) in the native state
do not have secondary structures accessible for external interactions. This is
prevented by the rigid nuclear structure of such proteins ([7], Lecture 13).
Thus, we see that the hypothesis of native aggregation differs from the
model accepted in the literature in that it involves nothing random and
nonspecific. Moreover, it contains elements of control and management: genetic
control of the primary sequence (hence, also the properties of secondary
structures), ligands, highly specific interactions of secondary structures with
each other, and spatial control of the course of native aggregation.
As for spatial control, it is also provided first of all by interactions of
"residual" secondary structures of neighboring natively unfolded
proteins (from the point of view of the proposed approach). This is quite a
substantive suggestion, if we take into account that the complete absence of
secondary structures is possible under the most severe conditions ([7], Lecture
17). If we also take into account the selective binding role of "residual"
secondary structures, the spatial structure of the protein matrix in the resting state is also
under genetic control, as properties of the "residual" secondary
structures are encoded by the primary sequence of amino acid residues.
Now let us consider the properties of a molten globule in greater detail. Packing of polypeptide chain of
normal globule is so dense that the side chains are tightly
apposed to each other and their rotation around valence bonds (turn
isomerization) is impossible. When the nucleus melts, the globules increase in
volume by approximately 50% [36]; free volume appears and, concomitantly, turn
isomerization also becomes possible. As a result of nuclear loosening, water
and hydrophobic substances (for instance, the dye ANS) begin to penetrate into
the nucleus. If the intensity of the denaturing factor rises, the molten
globule is converted into a premolten globule, in which the amount of secondary
structure is approximately half that in the molten globule ([7], Lecture 18).
These properties of the molten globule (to say nothing about the premolten one)
indicate that its nucleus loses rigidity and more closely resembles a fluid. An
elevation of conformational temperature inevitably leads to increased mobility
of parts of the molecule and to a decrease of the portion of the polypeptide
chain included in secondary structures. Modification of secondary structures
inevitably leads to a change of their specificity due to a change of their
topological characteristics. In other words, a change in size of secondary
structure (for instance, length of α-helix)
means a change in the biological meaning of the polypeptide
"sentence". The logic of this statement has been confirmed
experimentally in studies indicating that the nucleus of a molten globule is
structurally labile [40]. Thus, the molten globule is converted into a
reaction-capable protein that can participate in native aggregation.
Next, let us consider data indicating the involvement of the protein secondary
structures in mechanisms of signal transmission. Kim et al. [41] studied the
dynamics of the cytoplasmic domains of the E. coli chemotaxis receptor on
interaction with repellent and attractant. These authors concluded that an
attractant decreases the number of secondary structures in the domain, which
blocks signal transmission into the cytoplasm. A repellent produces the
opposite effect: it increases the amount of secondary structures in the domain,
and this makes the signal function of the receptor possible. In terms of the hypothesis
of native aggregation, repellent converts the domain into the excited state,
when its "valence" for interactions necessary for signal transmission
appears. The authors also believe that methylation/demethylation of receptors
is so important for their clustering and the dissociation of the formed
clusters because it causes significant changes in the amount of secondary
structures in domains.
Williams et al. [42] note that the orderliness of a polypeptide chain is
closely connected with protein function. Thus, for instance, binding of ligands
to streptavidin, purine nucleoside phosphorylase, hypoxanthine-guanine
phosphoribosyl transferase, hemoglobin, and myoglobin leads to some
disorderliness in the protein molecules. The authors performed thermodynamic
analyses of the actions of agonists and antagonists on the corresponding
receptors and came to the conclusion that mechanism of action of these ligands
was connected to opposite effects on the orderliness of the receptor structure;
denser polypeptide chain packing inside the protein leads to enhancement of the
degree of receptor oligomerization, while less dense packing decreases the
degree of oligomerization. Interestingly, agonists produce opposite structural
changes in different receptors. Thus, while an agonist of receptor 1 increases
polypeptide chain packing in receptor 1, an agonist for receptor 2 decreases
the packing in receptor 2. The same principle applies to antagonists. The
physiological sense of these changes will be understood only when it becomes
clear which part of which signaling pathway these changes constitute. Receptors
are one more system for which the existence of two states - resting and
activated - seems obvious. In this sense, the cell may be considered a
megareceptor: conversion from one state into another produces a complex signal
to neighboring cells.
According to current concepts, chaperones play an important role in cell life.
An example of interest is the small heat-shock proteins, a variable class of
chaperones widely distributed in cells of various types. Some representatives
of this family are inactive in cells in the resting state and are activated,
for instance, on heating [43]. According to the logic of native aggregation,
the triggering action of heat not only leads to the appearance of non-native
protein forms, but also activates the heat-shock proteins themselves. For this,
they must necessarily have either natively unfolded polypeptide chain regions
or the ability to be converted into the molten globule state. This will lead to
formation of a native aggregation center and then to aggregation itself. Native
aggregation of an activated
chaperone with an activated
target begins.
The presence of natively unfolded regions in chaperone molecules has been
accepted in the literature as necessary for their work [44-46]. From the point
of view of native aggregation, these unfolded sites are needed for the
formation of the secondary structures necessary for native aggregation with a
target. But the target itself is an excited protein that can become either a
natively unfolded protein (or have natively unfolded protein regions) or a
globule that becomes a molten globule. This suggestion is confirmed by the
studies of Hegyi and Tompa [47], who showed that natively unfolded proteins
have no tendency to interact with chaperones. This result is understood.
Natively unfolded proteins are proteins in the resting state. To interact with
other proteins, including chaperones, they must be activated - to be converted
into the excited, denatured state. On the other hand, chaperones have long been
known to be able to interact with molten globules [48].
From the point of view of the proposed approach, the results of native
aggregation are new structures necessary for the excited cell. The formation of
such structures is a cooperative process that needs the participation of two or
more proteins. Without such cooperation, the new structure cannot be created.
With such an understanding of native aggregation, it becomes obvious that each
of the two or more proteins, when interacting with each other, helps the
correct folding of the protein-partner. In other words, all proteins
participating in native aggregation are chaperones for each other, but some of
them might be more profoundly specialized in this direction.
It is well known that for the release of protein-targets, some chaperones need
ATP [43]. This fact is well explained in terms of Ling's concept: binding of
ATP leads to disassembly of the secondary structures formed in the natively unfolded
regions of the chaperone molecules bound to the protein-target. As a result,
the complex of chaperone with target is split. For other chaperones, the role
of ATP can be played by different ligands.
Native aggregation, like any other process in the cell, can be an object of
regulation. Its course can be affected by various factors that produce new
signaling structures. As an example, programmed cell death can be considered.
The mechanism of genetic regulation of apoptosis can be a source of the signal
that leads, as a result of native aggregation, to the appearance of a structure
that will trigger the whole cascade of reactions necessary for cell
degradation. From the point of view of native aggregation, such a structure can
appear in any part of cell - in the nucleus, cytoplasm, organelles, or plasma
membrane. The "structure of death" produced by native aggregation can
also appear if the cell (or any of its parts) is damaged. By the same
mechanism, other cell pathologies, for instance cancer, can appear.
With reference to the peculiarities of cancer cells, I would like to note one
feature that is directly related to the subject of the present paper. The
content of bound water in cancer cells is known to be lower than in precursor
cells [49]. It is on the basis of this difference that the technology of
magnetic resonance imaging allows malignant tumors to be recognized
non-invasively. From Ling's point of view, this means there are fewer natively
unfolded proteins in the cancer cell than in the normal cell. At the same time,
it has been shown in silico that the natively unfolded regions are more
extensive in cancer cell proteins: in cancer-associated proteins, the number of
such areas is 70% greater, while in signaling proteins it is 5 times greater [50].
It is obvious that natively unfolded proteins represent a diverse population
and are directly involved in cell transformations of pathological character.
Dynamics
of the hydrophobic phase of the living cell. As I already
mentioned, the cell in the resting state is a hydrophilic system. This is
confirmed by data on the distribution of hydrophobic substances (vital dyes)
between cell and medium under
conditions of steady-state distribution: the cell in the resting
state does not adsorb such substances [1].
Here I would like to draw the reader's attention to a very important
circumstance: under conditions of diffusional equilibrium, the plasma membrane
stops working as a barrier to a diffusing substance. There are no absolutely
impermeable membranes, especially for hydrophobic substances. The dye
undoubtedly penetrates into the resting cell, but is not accumulated in it.
Why? There are two reasons: in the cell, hydrophobic binding centers for dyes
are absent; and intracellular water is a poor solvent for them. For these
reasons, the dye molecules penetrating into the cell are eventually pushed out
into the medium. Thereby, under conditions of steady-state distribution, the character of
the distribution of the substance between cell and medium is determined by only
two factors: sorption on intracellular structures, and the low solving
capability of intracellular water ([51], Ch. 5).
Everything changes when the cell is converted to the excited state: binding of
vital dyes under conditions of steady-state distribution rises by tens or
hundreds percent of times [1]. Only one explanation for this is possible: the
volume of the hydrophobic phase in the cell increases explosively [8].
The hydrophobic phase is habitually associated with the membrane lipid phase;
but the volume of the lipid phase is negligible compared with cell size and,
what is most important, it cannot rise tens of times in fractions of second.
However, as we already know, proteins in the excited cell undergo
denaturational changes [1]. Hence, the cause of the increase of cell
hydrophobicity should be looked for in proteins rather than in lipids [8].
The hypothesis of native aggregation provides a simple explanation for
the hydrophobic burst in the cell: the cause of the increase of the hydrophobic
phase is the appearance of excited proteins. Indeed, according to the proposed
approach, when the threshold of perturbating action on natively unfolded
proteins is exceeded, secondary structures begin to form. These structures, in
the course of native aggregation, are then included in the hydrophobic areas of
new structures - structures of excitation. As stated earlier, the hydrophobic
areas are formed not only by molten globules, but also by the secondary
structures appearing on the folding of unfolded protein regions.
The high rate of formation of secondary structures, within the microsecond time
range ([7], Lecture 9), also determines the high rate of native aggregation
overall, which explains the hydrophobic burst in the excited cell. On the reverse
transition to the resting state, the cell again becomes hydrophilic. According
to the hypothesis of native aggregation, significant changes in the hydrophobic
phase can take place in any cellular structure, including membranes and
organelles.
The proposed existence of temporary hydrophobic protein phases explains
interesting phenomena known from pharmacology, when the efficiency of a
therapeutic agent depends on the degree of functional activity of the target
cell. The best known example of this seems to be verapamil. This hydrophobic
compound [52] scarcely affects the normal heart rhythm, but very efficiently
inhibits tachycardia. The same regularity is also observed in the action of
verapamil on skeletal muscle. This dependence can be explained if, on
excitation, verapamil-binding hydrophobic receptors appear in the muscle fiber.
The effect of verapamil is due to its blocking action on slow calcium channels;
but from the point of view of the principles of native aggregation, the cell
can also contain other dynamic hydrophobic targets for pharmacological agents
of various types. In other words, using the native aggregation principles, it
is possible to predict the existence of drugs acting only on the active cell;
their targets can be located not only in the membrane (as in the case of
verapamil), but also in other parts of the cell. Such medications will produce
no marked effect on cells in the resting state (the healthy state).
The role of the dynamic hydrophobic protein phase in the life of the cell has
not been studied at all. It is unknown in the equations of cell physiology. At
present, one can discuss the significance of this X-factor only in terms of
very general regularities based on simple physical principles. For instance, it
is obvious that the appearance of the hydrophobic phase in a cell will cause
the redistribution of all hydrophobic compounds including ATP [8]. The
redistribution of hydrophobic substances between the cell and the medium will
also begin.
However, the redistribution of substances is triggered not only by the
appearance of the temporary hydrophobic phase, but also by the desorption of
water from protein surfaces. As secondary structures start to form, the
adsorbed water will become free and the "bad" solvent will become
"good". This will lead to a rapid invasion of small solute molecules
into the areas that were previously occupied by adsorbed water. If we take into
account the rapid rate of formation of secondary structures ([7], Lecture 9),
it becomes obvious that during the fast destruction of the ordered water
structure, sharp concentration gradients of such substances will appear. In the
case of ions, everywhere in the cell, in microvolumes, significant diffusional
potentials will appear that may prove to be one cause of the appearance of
molten globules. Significant concentration gradients of dissolved substances
can also appear when the ordered water layers are restored, as the rate of
their restoration will also be determined by the high rate of disassembly of secondary
structures in activated proteins.
It is obvious that during the course of native aggregation the density and
rigidity of the protein matrix will increase owing to a rise in the number of
interprotein contacts. This provides even more difficulties for models of cell
function regulation that base their mechanisms on the free diffusion of
substances in the cell, since with an increase of protein matrix density the
significance of diffusional processes will decrease.
If we return to the cell protoreaction, it can be concluded with certainty that
the hypothesis of native aggregation has managed to explain the rise of
viscosity and turbidity of the cytoplasm (Fig. 1) as well as the increase of
volume of the cell hydrophobic phase. From the proposed mechanism it is clear
that the changes discussed will occur synchronously, as the key link among all
these changes is the structural readjustment of the same key proteins.
Conclusion. The cornerstone of the hypothesis of native aggregation is the generation in proteins of temporary secondary structures that can interact selectively with secondary structures in the same or other proteins. The nonspecific reaction of cells, which was studied by Nasonov's school, turns out in reality to comprise myriads of specific protein-protein interactions. Since native aggregation is directed by active secondary protein structures, it proves to be completely under genetic control, so the dogma of Anfinsen [53] formulated for the folded polypeptide chain can be extended by incorporating native aggregation into its sphere of application.
Acknowledgments. I am very grateful to Paul Agutter, James Clegg, Ilya Digel, Laurent Jaeken, José Neira and Richard Wiggins for valuable critical comments on this article. I also appreciate Leonid Pevzner's assistance in preparation of this paper.
References
1. Nasonov DN:
Local
Reaction of Protoplasm and Gradual Excitation (English Transl. by Halpern,
Y.S.), Washington, D.C.: National Science Foundation, available at Office of
Technical Services, US. Department of Commerce; 1962.
2. Ling GN:
A
Physical Theory of the Living State: The Association-Induction Hypothesis.
Waltham MA: Blaisdell; 1962.
3. Ling GN:
In
Search of the Physical Basis of Life. New York: Plenum Publ Corp; 1984.
4. Ling GN:
A
Revolution in the Physiology of the Living Cell. Malabar FL: Krieger Publ
Co; 1992.
5. Ling GN:
Life
at the Cell and Below-Cell Level: the Hidden History of a Fundamental
Revolution in Biology. New York: Pacific Press: 2001. (The Russian
translation was published in 2008).
6. Ling GN:
A
convergence of experimental and theoretical breakthroughs affirms the PM theory
of dynamically structured cell water at the theory's 40th birthday. In
Water and the Cell. Edited by Pollack GH, Cameron IL and Wheatley DN. Berlin,
New York: Springer Verlag; 2006:1-52.
7. Finkelshtein AV, Ptitsyn OB: Physics of Proteins. Amsterdam: Academic Press;
2002.
8. Matveev VV: Protoreaction
of protoplasm. Cell Mol Biol 2005, 51:715-723.
9. Wilson, EB: The Cell in Development and Heredity. New York: Macmillan; 1928.
10. Porter KR: The
cytomatrix: a short history of its study. J Cell Biol 1984, 99:3s-12s.
11. Porter KR, Tucker JB: The ground substance of the living cell. Sci Am 1981,
244:56-67.
12. Heuser J: Whatever happened to the 'microtrabecular concept'? Biol Cell
2002, 94:561-596.
13. Fulton AB: How crowded is the cytoplasm? Cell 1982, 30:345-347.
14. Baló-Banga
JM, Molnár
L, Nováki M,
Leibinger J: Measurement
of lymphocyte activation by a chromatin topo-optical reaction. Mechanism and
specificity of the test. Arch Dermatol Res 1980, 269:239-251.
15. Baló-Banga
JM, Molnár
L, Nováki M,
Leibinger J: Measurement of lymphocyte activation by a chromatin topooptical
reaction. II. Application for detecting drug allergy. A clinical and
experimental study. Allerg Immunol (Leipz) 1980, 26:137-153.
16. Inners D, Bendet IJ: Thermal stability of T2 DNA in situ. Virology 1969,
38:269-277.
17. Bearden J Jr, Bendet IJ: Birefringence of spermatozoa. I. Birefringence melting
of squid, bull, and human sperm nucleoprotein. J Cell Biol 1972, 55:489-500.
18. Mirsky AE, Pauling L: On the Structure of Native, Denatured, and Coagulated
Proteins. Proc Natl Acad Sci USA 1936, 22:439-447.
19. Mirsky AE: Protein coagulation as a result of fertilization. Science 1936,
84:333-334.
20. Mirsky AE: The
Visual Cycle and Protein Denaturation. Proc Natl Acad Sci USA 1936, 22:147-149.
21. Proskuryakov SY, Konoplyannikov AG, Gabai VL: Necrosis:
a specific form of programmed cell death? Exp Cell Res 2003, 283:1-16.
22. Ling GN: The role of phosphate in the maintenance of the resting potential
and selective ionic accumulation in frog muscle cells. In Phosphorus metabolism
vol. II. Edited by McElroy WD and Glass B. Baltimore: The Johns Hopkins
University Press; 1952:748-795.
23. Ling GN: The
physical state of water in living cell and model systems. Ann NY Acad Sci
1965, 125:401-417.
24. Spackman MA, Munshi P, Dittrich B: Dipole
moment enhancement in molecular crystals from X-ray diffraction data.
Chemphyschem 2007, 8:2051-2063.
25. Kemp DD, Gordon MS: An
interpretation of the enhancement of the water dipole moment due to the
presence of other water molecules. J Phys Chem A 2008, 112:4885-4894.
26. Collins JM, Leadbeater NE: Microwave
energy: a versatile tool for the biosciences. Org Biomol Chem 2007,
5:1141-1150.
27. Ling GN: Nano-protoplasm:
the ultimate unit of life. Physiol Chem Phys Med NMR 2007, 39:111-234. See also: Ling GN, Ochsenfeld MM. A
historically significant study that at once disproves the membrane (pump)
theory and confirms that nano-protoplasm is the ultimate physical basis of
life--yet so simple and low-cost that it could easily be repeated in many high school biology classrooms worldwide. Physiol
Chem Phys Med NMR. 2008;40:89-113.
28. Zheng JM and Pollack GH: Long-range
forces extending from polymer-gel surfaces. Phys Rev E 2003, 68:031408.
29. Zheng JM, Chin WC, Khijniak E, Khijniak E Jr, Pollack GH: Surfaces
and interfacial water: evidence that hydrophilic surfaces have long-range
impact. Adv Colloid Interface Sci 2006, 127:19-27.
30. Ling GN: The physical state of water and ions in living cells and a new
theory of the energization of biological work performance by ATP. Mol Cell
Biochem 1977, 15:159-172.
31. Iakoucheva LM, Radivojac P, Brown CJ, O'Connor TR, Sikes JG, Obradovic Z,
Dunker AK: The importance of intrinsic disorder for protein phosphorylation.
Nucleic Acids Res 2004, 32:1037-1049.
32. Mukhopadhyay S, Krishnan R, Lemke EA, Lindquist S, Deniz AA: A natively
unfolded yeast prion monomer adopts an ensemble of collapsed and rapidly fluctuating
structures. Proc Natl Acad Sci USA 2007, 104:2649-2654.
33. Dunker AK, Lawson JD, Brown CJ, Williams RM, Romero P, Oh JS, Oldfield CJ,
Campen AM, Ratliff CM, Hipps KW, Ausio J, Nissen MS, Reeves R, Kang C,
Kissinger CR, Bailey RW, Griswold MD, Chiu W, Garner EC, Obradovic Z:
Intrinsically disordered protein. J Mol Graph Model 2001, 19:26-59.
34. Dunker AK, Obradovic Z, Romero P, Garner EC, Brown CJ: Intrinsic protein
disorder in complete genomes. Genome Inform Ser Workshop Genome Inform 2000, 11:161-171.
35. Dunker AK, Brown CJ, Lawson JD, Iakoucheva LM, Obradovic Z: Intrinsic
disorder and protein function. Biochemistry 2002, 41:6573-6582.
36. Uversky VN: Natively unfolded proteins: a point where biology waits for
physics. Protein Sci 2002, 11:739-756.
37. Eliezer D, Palmer AG 3rd: Proteins hunt and gather. Nature 2007,
447:920-921.
38. Wright PE, Dyson HJ: Linking
folding and binding. Curr Opin Struct Biol 2009, 19:31-38.
39. Ellis RJ: Macromolecular crowding: obvious but underappreciated. Trends
Biochem Sci 2001, 26:597-604.
40. Lala AK, Kaul P: Increased exposure of hydrophobic surface in molten
globule state of alpha-lactalbumin. Fluorescence and hydrophobic photolabeling
studies. J Biol Chem 1992, 267:19914-19918.
41. Kim SH, Wang W, Kim KK: Dynamic and clustering model of bacterial
chemotaxis receptors: structural basis for signaling and high sensitivity. Proc
Natl Acad Sci USA 2002, 99:11611-11615.
42. Williams DH, O'Brien DP, Sandercock AM, Stephens E: Order changes within
receptor systems upon ligand binding: receptor tightening/oligomerisation and
the interpretation of binding parameters. J Mol Biol 2004, 340:373-383.
43. Haslbeck M, Franzmann T, Weinfurtner D, Buchner J: Some like it hot: the
structure and function of small heat-shock proteins. Nat Struct Mol Biol 2005,
12:842-846.
44. Weikl T, Abelmann K, Buchner J: An
unstructured C-terminal region of the Hsp90 co-chaperone p23 is important for
its chaperone function. J Mol Biol 1999, 293:685-691.
45. Richardson A, Schwager F, Landry SJ, Georgopoulos C: The importance of a mobile
loop in regulating chaperonin/co-chaperonin interaction: humans versus
Escherichia coli. J Biol Chem 2001, 276:4981-4987.
46. Tompa P, Csermely P: The
role of structural disorder in the function of RNA and protein chaperones.
FASEB J 2004, 18:1169-1175.
47. Hegyi H, Tompa P: Intrinsically
disordered proteins display no preference for chaperone binding in vivo. PLoS Comput
Biol 2008, 4:e1000017.
48. Rawat U, Rao M: Interactions of chaperone alpha-crystallin with the
molten globule state of xylose reductase. Implications for reconstitution of
the active enzyme. J Biol Chem 1998, 273:9415-23.
49. Chaplin M: Do we underestimate the importance of water in cell biology? Nat
Rev Mol Cell Biol 2006, 7:861-866.
50. Iakoucheva LM, Brown CJ, Lawson JD, Obradovic Z, Dunker AK: Intrinsic
disorder in cell-signaling and cancer-associated proteins. J Mol Biol 2002,
323:573-584.
51. Troshin AS:
Problems
of cell permeability. London, New York: Pergamon Press, 1966.
Edition of 1956 in Chinese:
PDF
DJVU
52. Kasim NA, Whitehouse M, Ramachandran C, Bermejo M, Lennernäs H, Hussain AS,
Junginger HE, Stavchansky SA, Midha KK, Shah VP, Amidon GL: Molecular
properties of WHO essential drugs and provisional biopharmaceutical
classification. Mol Pharm 2004, 1:85-96.
53. Anfinsen CB: Principles that govern the folding of protein chains. Science
1973, 181:223-230.
Full
text [PDF]